Constant with this particular, when purified human fibrocytes have been intravenously transferred to SCID mice that had been exposed to both bleomycin or saline, higher numbers of human CD45 Col1 CXCR4 fibro cytes were observed in bleomycin challenged lungs as in comparison to saline treated controls. In examining the chemokine receptor profile of circulat ing CD45 Col1 cells in mice, we’ve found that the two in normal mice and animals challenged with bleomycin, CXCR4 would be the most generally expressed surface receptor, remaining present in roughly 70% from the cells. Additionally, CXCL12, the ligand for CXCR4, is expressed while in the lungs and is induced immediately after intrapulmonary administra tion of bleomycin inside 1 day and then stays ele vated for your subsequent 19 days, the dynamics of this expression consequently are constant that has a function for CXCL12 in recruiting CXCR4 fibroytes towards the lungs.
Indeed, in vivo neutralization of CXCL12 resulted in diminished quantity of lung CD45 Col1 CXCR4 fibro cytes and also a SMA expressing myofibroblasts selleck as well as decreased lung collagen written content and attenuated pulmonary fibrosis by histologic morphometric analysis, but did not influence the number of lung neutrophils, macrophages, CD4 and CD8 T cells or NK cells. Constant with this, pharmacological antagonism of CXCR4 also results in reduced lung fibrocyte numbers and pulmonary fibrosis in response to bleomycin. Operate by other groups has examined the part of other mechanisms in recruitment of fibrocytes to your lungs in animal designs of lung fibrosis.
Utilizing a model of intrapul monary fluorescein isothiocyanate induced lung fibrosis, fibrocytes have been isolated from lung tissue and bronchoal veolar lavage just after in vitro culture. These cells expressed CXCR4, CCR5, CCR7 and CCR2 and migrated in response to CCL2 and CCL12 ligands. CCR2 deficient mice treated with intratracheal investigate this site FITC were identified to get decrease levels of fibrocytes within the lung and less fibrosis as when compared with wildtype counterparts, and effect that was later observed to be independent of CCL2, but was attributed to another CCR2 ligand, CCL12. CCR2 can be very expressed on cells on the mononuclear phago cyte lineage like monocyte, macrophage and dendri tic cell populations, even so, and lowered lung fibrosis in response to bleomycin in CCR2 knockout animals corre lated using a substantial reduction in these cells also as inflammatory cytokines during the bronchoalveolar lavage fluid, it’s consequently not clear no matter whether the observed impact in CCR2 deficient animals is attributable to fibro cytes or other cell populations.
Fibrocyte influx on the lung inside the bleomycin model has also been linked to your CCL3 CCR5 chemokine axis, interestingly, this effect was asso ciated with reduced lung expression of lung CXCL12 expression from the lungs of CCL3 and CCR5 deficient ani mals, suggesting that the impact of CCL3 CCR5 may perhaps be mediated by way of the CXCL12 CXCR4 axis.
Monthly Archives: May 2014
Simulation study We developed and carried out a series of simul
Simulation review We made and carried out a series of simulations to additional assess our proposed technique. We applied the fitted model obtained from applying iBMA just before the yeast time series microarray data set since the genuine underlying network, and generated simulated expres sion information in the estimated linear regression model. Twenty information sets, every single with all the very same dimensions as the genuine time series expression data, were independ ently generated as follows, 1. Set the prior probability of the regulatory romantic relationship for every gene pair for the similar value as the regulatory probable obtained in the supervised finding out stage applying the actual external information. two. Set the expression amounts in the 3556 genes to the 95 yeast segregants and also the two parental strains at time t 0 as the observed measurements from the serious yeast time series gene expression data.
three. For kinase inhibitor TAK 165 each target gene g, define the set Rg of real regulators as those using a posterior probability of 50% in our inferred network applying iBMA prior as well as the authentic time series information. four. For time t one to 5, tematically integrates external biological know-how into BMA for network development. A key attribute of our ap proach is really a formal mechanism to account for model un certainty. For every target gene, we arrive at a compact the place the Bs are provided through the posterior expectation of the regression coefficients corresponding for the set of correct regulators determined in Step three. five. Generate the simulated observed gene expression ranges by adding noise on the real expression amounts with no measurement mistakes, i.
e, in which Eg,t,s N with ?two staying given from the sample variance on the regression residuals during the authentic information evaluation. Others, e. g, have proven that the error in log ratios of expression data is fairly around selleck inhibitor by a ordinary distribution. To assess the accuracy of networks inferred together with the simulated information sets, we in contrast each of those net performs for the correct network created in Step 3 of your data generation algorithm. We made use of the identical evaluation cri teria as during the authentic information evaluation together with the genuine network replacing  Yeastract as the reference. As shown in Table 5, iBMA prior out performed the other iBMA based mostly solutions, yielding a TPR of 71. 13% averaged over 20 replications. set of promising designs from which to draw inference, the weights of that are calibrated from the external bio logical knowledge. Our approach infers sparse, compact and accurate networks upon the input of the acceptable estimate of network density from each real and simu lated data. It does not place a hard restrict about the amount of regulators per target gene, in contrast to another approaches, this kind of as Bayesian network approaches that impose this constraint to cut back the computational burden.
Yeastract as the reference. As shown in Table 5, iBMA prior out performed the other iBMA based mostly solutions, yielding a TPR of 71. 13% averaged over 20 replications. set of promising designs from which to draw inference, the weights of that are calibrated from the external bio logical knowledge. Our approach infers sparse, compact and accurate networks upon the input of the acceptable estimate of network density from each real and simu lated data. It does not place a hard restrict about the amount of regulators per target gene, in contrast to another approaches, this kind of as Bayesian network approaches that impose this constraint to cut back the computational burden.
First, 16S rRNA gene sequences have been retrieved and compared t
Initially, 16S rRNA gene sequences had been retrieved and compared to a database of regarded 16S rRNA gene sequences. Every read through that matched a recognized sequence was assigned to that organism. In the second analysis putative open studying frames have been identified and their corresponding protein sequences were searched with BLAST against the M5NR database. The M5NR is surely an integration of a lot of sequence databases into one single, searchable database. This approach professional vided us with info for assignments to taxonomic units using the caveat a protein sequence could possibly be assigned to greater than one particular closely related organism. Taxonomic assignments were resolved using the lowest common ancestor ap proach. Practical examination and reconstruction of metabolic pathways ORFs have been identified and their corresponding protein sequences have been annotated by comparison to SEED, Pfam, TIGRfam and COG information bases.
Recognized proteins were assigned with their respective enzyme commission number. Just before quantitative characterization, counts had been normalized towards the complete quantity of hits inside their respective database applying successful sequence counts, a composite order AZD2171 measure of sequence variety and average genome size from the metagenome as described by Beszteri et al. Raes and colleagues defined the AGS as an ecological measure of genome dimension that also includes a number of plas mid copies, inserted sequences, and connected phages and viruses. Previous studies demonstrated the relative abundance of genes will show variations if the AGS on the local community fluctuate across samples. The ChaoI and ACE estimators of COG richness had been computed together with the software SPADE v2.one utilizing the number of person COGs per unique COG function. The proportion of spe cific genes in metagenomes also gives a approach for comparison amongst samples.
By dividing the AGS to your amount of DNA per function certain gene, a single can determine the proportion of genomes during the metagenome which can be capable of that perform. However, direct comparison from the distribution selleck chemical of vary ent functions was not established involving the metagenome, considering that length and copy number of the gene was not incorporated in the formula. To define whether a gene was enriched within the natural environment we calculated the odds ratio or the relative risk of observing a offered group in the sample relative to your comparison dataset. The odds ratios were calculated as follows, where A is the amount of hits to a offered category while in the x dataset, B would be the number of hits to all other categories inside the x metagenome, C will be the number of hits to a given class within the y dataset, and D will be the amount of hits to all other classes inside the y dataset. We then utilised the metagenome profiles to determine the statistical differ ences involving the two samples based mostly around the Fishers precise test with corrected q values using the software package deal STAMP v1.
Open go through frames in just about every transcriptome assembly
Open go through frames in every transcriptome assembly had been searched using scripts presented from the TRINITY pipeline. The TRINITY system essentially implements the ORF prediction techniques of GENEID, We searched for that 500 lon gest ORFs in all 6 reading frames in each dataset and utilised these to parameterize a hexamer primarily based Markov model. Exactly the same ORFs were then randomized to create a null model for non coding sequence and all transcripts had been then searched for that longest, probably coding ORF. This was scored as putatively coding or non coding in accordance to a probability ratio check. Clusters of gene families were developed applying the predicted proteins of T. californicum, T. grallator and picked outgroups with fully sequenced genomes. If iso types for a gene existed from the predicted peptides of the Theridion species, only the longest variant was retained.
For outgroup comparisons, essentially the most latest CDS se quences were selected through the following taxa with current genome sequences. Nematostella vectensis, Homo sapi ens, Daphnia pulex, Nasonia vitripen nis, Tro bolium castaneum, Drosophila melanogaster, and Tetra nychus urticae, annotation stage and don’t seem in Figure two. Phylogenetic inference Orthologous EVP4593 ic50 genes have been identified utilizing the HAMSTR pipeline, HAMSTR utilizes hidden Markov versions and reciprocal very best hit BLAST searches against a predefined set of orthologous sequences de rived from model organisms. The recognized orthologs had been aligned individually. The plans GBLOCKS, ALISCORE, and ALICUT had been made use of to eliminate poorly aligned and overly gappy portions of the alignments. Sequences under one hundred amino acids in length had been eliminated, and any alignments with missing taxa had been deleted.
The 352 trimmed alignments remaining, comprising 170,965 aligned amino acid internet sites, have been concatenated using FASconCAT, as well as a parti tioned highest likelihood phylogenetic analysis run from the program RAXML, The concatenated alignments were partitioned by selleck tsa inhibitor gene, and every single partition was assigned the PROTGAMMA model utilizing the WAG amino acid substitution matrix, To seek out essentially the most probably tree topology, one thousand random addition sequence replicates had been carried out followed by one thousand bootstrap replicates. The chronopl command through the R bundle APE was utilized to produce an ultrametric phylogeny via the non parametric rate smoothing technique using the RAXML tree. The evaluation applied no fossil or other calibration points, so the branch lengths show time in evolutionary units from 0 to 1.
The gut represents the first line of defense against ingested hos
The gut represents the 1st line of defense towards ingested host plant allelochemicals, pesticides, and other toxins and lots of transcripts predicted to encode detoxi fication enzymes have been detected. One example is, 50 cyto chrome P450 like unigenes have been detected inside the A. glabripennis midgut transcriptome. These enzymes have versatile oxidoreductive properties, are very involved with degrading lipophilic harmful toxins, and also have been proven to confer resistance to pesticides as well as little aromatic toxins that could accumulate to large con centrations while in the heartwood of trees, Nearly all the cytochrome P450 unigenes detected while in the A.
glabripennis midgut transcriptome had highest scoring BLASTP alignments to cytochrome P450s iden tified within the Tribolium castaneum genome, though the % similarity with the amino acid level ranged from 34% to 66%, reflecting a comparatively substantial degree of Ivacaftor structure divergence from previously annotated cyto chrome P450s, Cytochrome P450 unigenes detected while in the A. glab ripennis midgut had been putatively assigned to clans and families dependant on the annotations in the highest scoring BLASTP alignments, To conform to cytochrome P450 classification conventions, only alignments sharing 40% amino acid identity were used to annotate these unigenes. Even though a handful of unigenes predicted to encode mitochondrial cyto chrome P450s had been recognized, non mitochondrial clans three and 4 have been extra hugely represented in com parison.
Non mitochondrial cytochrome P450s had been classified to five families, which have been predominated by CYP6, but also included CYP4, CYP9, CYP345, and CYP347, CYP6 relatives genes are often existing in numerous copies, selleck ABT-263 arise in clusters in insect genomes, have pivotal roles during the detoxi fication of host plant defensive chemical substances this kind of as xanthotoxins, gossypol, and chlorogenic acid, and are generally induced in herbivorous insects through periods of feeding. Unigenes predicted to encode carboxylesterases have been the most dominant enzyme with putative involvement in detoxification processes detected within the midgut. 65 individual unigenes had been predicted to encode these proteins. While secreted carboxylesterases are gener ally involved with pheromone metabolism, intracellular carboxylesterases are often implicated in pesticide and allelochemical metabolism and tolerance, In excess of 30 of your carboxylesterase unigenes detected within the A. glabri pennis midgut lacked secretory peptides and may very well be primed to serve detoxification roles. For example, automobile boxylesterases are hypothesized to mediate resistance to phenolic glycosides in Papilio canadensis, and therefore are usually uncovered in substantial amounts from the midgut, Notably, trees during the household Salicaceae, which incorporate a lot of A.
gambiae was most likely a refinement of that of An quadriannulat
gambiae was more than likely a refinement of that of An. quadriannulatus. Moreover, this biased receptivity of An. gambiae antenna towards human derived odors may very well be additional augmented by the practical variations involving orthologous ORs advised by our sequence analyses. Potential practical exams of AqOr odor tuning will more increase our understanding within this regard. Taken with each other, and provided the central position that ORs perform in defining host specificity, the anthropophagy of An. gambiae is almost certainly not derived from the evolution of any single OR certain for your function of human host in search of. Alternatively, we posit the receptivity bias in the antenna of An. gambiae towards human host odors is probable the consequence from the cumulative results of the two practical divergences and modifications during the abundance and distribution of typical ORs currently existing inside the An.
gambiae species complicated. Approaches Gene annotation The genome assemblies of An. gambiae and An. quadriannulatus had been downloaded from the sites of VectorBase and Broad Institute, respectively. The annotation of chemosensory genes was carried out following a past protocol, In brief, previously reported selleckchem Amuvatinib chemosensory genes from An. gambiae, Aedes aegypti, Culex quinquefasciatus, and D. melanogaster have been employed as queries in TBLASTN searches towards the two anopheline genomes. Putative chemosensory gene coding loci were identified following filtering out lower scoring blast hits. For every locus, the query sequence that yield the highest bit score was chosen as reference to perform homology based gene prediction working with GeneWise, All gene designs were manually inspected and modified if required.
All genomic information is available by means of VectorBase along with the annotated chemoreceptor sequences are listed in supplementary Table S1. Phylogenetic analysis For every of the OR GR IR OBP families, protein sequences of genes in the two mosquitoes were aligned employing MAFFT, The many sequence alignments have been manually curated and poorly aligned regions have been eliminated using trimAl with automated1 option. selleck chemicals TW-37 Maximum probability trees had been constructed working with RAxML along with the reliability of tree topology was evaluated with a hundred bootstrap replicates. Resulting gene trees have been recon ciled using the species phylogeny to estimate ancestral gene copy numbers and gene attain and loss events. An orthologous group is defined as a really supported clade representing a single gene while in the widespread ancestor of An.
gambiae and An. quadriannulatus. Analysis of sequence divergence For each orthologous pair of chemosensory genes in An. gambiae and An. quadriannulatus, protein sequences had been aligned using MAFFT as well as the corresponding nucleotide alignment was generated employing a customized script, 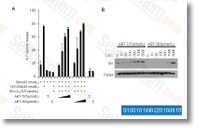 The price of amino acid substitution and dN dS ratio have been calculated using PROTDIST and CodeML, respectively.
The price of amino acid substitution and dN dS ratio have been calculated using PROTDIST and CodeML, respectively.
We then cross refer enced this for the down regulated gene set in
We then cross refer enced this to the down regulated gene set in handle versus muscle significantly less humeri, noting any genes enriched in excess of 3 fold in mesenchyme compared to regulate humeri. these are indicated in column two of Additional file one. Table S2. It can be feasible that these genes are involved in the two cartilage and muscle development so no genes happen to be eliminated through the data set, even so, DE genes also showing increased expression in mesenchyme com pared to regulate humeri will have to be taken care of with caution with respect to a skeletal certain response to mechanical stimu lation. Such genes have not been prioritised in any of our subsequent exploration of candidate mechanosensitive genes. The developing humerus at TS23 constitutes various cell and tissue populations at unique stages of differen tiation which includes the joint region, the perichondrium along with the organised zones inside the cartilage rudiment.
Therefore the experimental design and style employed right here will capture genes linked with unique cells styles at dif ferent stages of differentiation. It is going to now be crucial that you type out which cells and tissues have altered expres sion of exact genes. This could be selleck DOT1L inhibitor addressed for a sub set of genes by in situ hybridisation, with an original ana lysis of four genes presented in Figure 6. It may possibly be ad dressed in the high throughput method by isolating distinct cell populations working with laser microdissection from tissue sections, purification of RNA and quantitative RT PCR gene expression profiling, evaluating handle and mutant tissue from, one example is the hypertrophic, prehypertrophic or the elbow joint re gion alone.
We employed each RNA sequencing and Micro array technologies in parallel to determine dif ferential expression. Microarray technologies continues to be utilised to find out expression of chondrogenic and osteogenic selleck inhibitor genes from establishing total tissues, and from in vitro differentiation procedures, The usage of RNA seq technology to de scribe the transcriptome is a lot more recent, Previ ous direct comparisons concerning microarray and RNA sequencing primarily based approaches to reveal alterations in gene expression amongst tissues reported that RNA seq recognized far more DE genes, We also observed that RNA seq is even more delicate in reproducibly detecting alterations in gene expression, detecting far more genes al tered at reduced quantitative levels, This was additional emphasised by minimizing the stringency within the statistical analysis to p 0.
08, which enhanced the number of genes detected by micro array especially, An example on the im portance within the improved sensitivity and reproducibility of RNA seq is proven from the Spp1 gene which didn’t present statistical significance by microarray but has been verified by qRT PCR and in situ hybridisation, The more substantial dynamic array and higher reproducibility across replicates has also been noticed in other studies.
A pharmacophore conveys minimum traits on the structures of your
A pharmacophore conveys minimum traits of your structures from the ligands that are vital for binding on the target. Every hypothesis is accompanied using a set of aligned confor mations that propose the mode through which molecules are probable to bind comparatively, Following choosing the maxi mum quantity of web sites to be five, alignments were created utilizing model ligands which have been a part of the active set in the series. The compounds A30, A35 and B14 were not chosen out of complete 29 for alignment as a result lowering the number of matches to 26 for each of the hypotheses. The hypotheses obtained in addition to their survival scores and selectivity are reported in Table 3. Scoring perform So as to assign a score, every pharmacophore in addition to its ligand are temporarily regarded as the reference and various non reference pharmacophores are aligned 1 by one particular using a least square procedure.
Even further a web site score, a vector score and a volume score are calculated with combined weights for every aligned pharmacophore. Pharmacophoric hypotheses had been scored within the basis of how great the alignment exists amongst the energetic set molecules and selelck kinase inhibitor pharmacophoric benefits. Right after picking out the hypothesis for every box, the ultimate scoring is done and the resultant is known as the survival score of the hypothesis that characterizes its validity and probable to be utilized for a provided set of molecules. The survival score constitutes a number of different scores and weights calculated all through hypothesis generation as presented in equation 2.
where Ws will be the weights and Ss would be the scores We chosen a prevalent pharmacophore hypothesis comprising of prevalent chemical capabilities in the aligned active molecules in the congeneric set. The ultimate hypothesis, DDHRR. 8 was chosen primarily based on large selectivity at the same time because the survival score Amuvatinib structure which yields the perfect align ment from the active set ligands. Coupled with the internet site score, vector score and volume score DDHRR. eight was the perfect selection for browsing a compound library. Plainly, the name DDHRR implies the presence of two hydrogen donors, 1 hydrophobic group and two aro matic rings. In Figure five the hydrogen donors are marked with light blue spheres centered about the H atom with the arrows directing in direction of potential H bonds and the aro matic rings marked as group web-sites represented by orange torus, situated on the centroid of a group of atoms.
These marked internet sites give an strategy regarding the mode of inter action of the lead molecule with Cathepsin L. As repre sented in Figure five, an alignment within the 26 compounds through the congeneric series with that from the picked hypothesis, DDHRR. 8, supported its selection since the typical pharmacophore hypothesis. In Figure 5, intersite distances are shown. Very similar hypotheses were  grouped with each other according to their intersite distances to identify the common pharmacophore.
grouped with each other according to their intersite distances to identify the common pharmacophore.
Inhibitors for ERK1 2 and its con trol substance, p38, PI3K, Akt
Inhibitors for ERK1 two and its con trol substance, p38, PI3K, Akt were from Calbiochem, Rabbit polyclonal p38 MAPK and Akt, and monoclonal phos pho p38 MAPK, phospho Akt, p44 p42 MAPK, and phospho p44 p42 MAPK antibodies were obtained from Cell Signaling Technological innovation, Inc, Mouse monoclonal GAPDH, employed as western blot handle for equal gel load of protein, was bought from Santa Cruz Biotechnol ogy Inc, As assessed by the Limulus Amebocyte Lysate Pyrotell, the thrombin and trypsin preparations in our review contained under 0. 03 EU LPS ml. RNA isolation, reverse transcription and Quantitative RT PCR Single stranded cDNA was synthesized from total RNA and made use of to complete QRT PCR with gene particular pri mers as described previously, Unstimulated HOKs and samples without having RT served as detrimental controls.
A melting curve was performed with the finish of each QRT PCR to make certain the gene merchandise was specific. Gly ceraldeyde three phosphate selleck chemical dehydrogenase was used since the picked housekeeping gene for normaliza tion. The primer sequences for GAPDH, CXCL5, CXCL3 and CCL20 have already been described previously, Information were analyzed applying Pfaffl technique to cal culate fold alter of gene expression, Measurement of chemokines in culture supernatant Secreted CXCL5 and CCL20 have been measured in culture supernatant using Duoset ELISA kit, The absolute concentration in each and every sample was calculated depending on sample yielding optical density making use of the common curve. Chemokine secretion from just about every situation was normalized to chemokine developed at baseline level in unstimulated handle group, and results are presented as % of unstimulated manage cells.
ELISA assay for quantification of p38 and ERK1 two phosphorylation To assess phosphorylation of p38 and ERK1 2, handled cells had been homogenized in lysis buffer in accordance together with the companies protocol, Instantly before including to cells, protease and phosphatase inhi bitors, PMFS and sodium orthovanadate, Halt Protease Inhibitor Cocktail and Halt selleck inhibitor Phosphatase Inhibitor Cocktail, had been additional to your lysis buffer. Phosphorylation of p38 and p42 44 was assessed in cell lysate using Duoset antibody pairs following manufacturers protocol, Complete p38 was quantified by a very similar process and served for normalization. Total p38 was proportional to total protein measured in each and every sam ple and to the quantity of cells plated in each very well.
Western blot evaluation Total and phosphorylated kinase expression in untreated and agonist stimulated HOK cells was determined by Western immunoblot examination. Samples of entire cell lysate supernatant proteins, solubilized in 1X NuPage LSD sample and decreasing buffer, had been resolved conco mitantly with Precision Plus Dual Colour protein molecu lar weight normal in NuPAGE four 12% Bis Tris mini gel and MOPS SDS working buffer in XCell SureLock Mini Cell, trans ferred to PVDF membrane in XCell II Blot Module making use of electrophor esis techniques and protocols of InVitrogen Corp, PVDF membranes have been blocked for 1 h with 1% BSA, 1%PVP and 1% PEG in wash buffer, then.
GeneCoDis3 was then employed to uncover enriched biological proce
GeneCoDis3 was then employed to search out enriched biological method gene ontology terms and gene annotation co occurrence clusters that had been drastically connected together with the differentially expressed orthologues, In addition to single BP GO annotations, annotation clusters repre sented considerable relationships amid biological course of action, molecular perform, InterPro protein domain, and KEGG pathway annotations that were associated together with the vary entially expressed orthologoues, A reference back ground checklist of all D. melanogaster genes orthologous to all A. mellifera RNAs was utilised to find out significant en richment of BP GO terms and annotation clusters applying a chi square test corrected for several testing, Sig nificantly enriched BP GO terms that contained only one orthologue had been discarded.
The annotated transcripts and orthologues that exhibited differential expression had been compared. To deal with the hypothesis that diet program and early grownup improvement inhibitor HDAC Inhibitor impact A. mellifera gene expression and physiology, annotated transcripts that improved or decreased as a result of starvation or as folks aged from 3d to 8d old have been compared and biological processes and annotation clusters corresponding to differentially expressed orthologues have been analyzed for sizeable en richment. To tackle the hypothesis that starvation af fects younger and older workers in a different way, comparisons had been produced amongst annotated transcripts and ortholo gues that have been affected by diet program at either 3d or 8d of age. Venn Diagrams were constructed working with the annotated transcripts that have been differentially expressed due to eating plan at 3d and those that have been differentially expressed due to diet program at 8d.
To tackle the last hypothesis that create ment and aging are affected by eating plan, comparisons had been made involving annotated transcripts and orthologues that transformed with age for bees fed either a wealthy diet plan or individuals fed a bad food plan. Venn Diagrams have been constructed utilizing the annotated transcripts selleckchem that had been differentially expressed because of age for that rich eating plan and those who were differentially expressed as a result of age in bees deprived of pollen. rt PCR validation of gene expression modifications resulting from eating plan To assess the predictive electrical power with the mRNA seq analyses, we measured the expression of 6 genes vermiform, cdc42, E75, GTP binding protein ten, vitellogenin, glucocerebrosidase transcript variant 1 that had been signi ficantly differentially expressed based on the preceding mRNA seq experiment.
Body fat bodies of 8d previous bees from 3 replicate host colonies have been utilised. These bees were not utilised within the former mRNA seq experiment, however the colonies they came from shared related population sizes, age distributions, and distribution of food outlets on the 3 colonies used from the mRNA seq experiments and also to each and every other. Dietary manipulations, body fat entire body dissec tions, and RNA isolations were as described above except RNA integrity was not assayed.
